Ethan Bier
Research
There are two primary areas of research in my lab: 1) developmental patterning, and 2) using Drosophila as model to understand human disease processes.
Developmental Studies
With respect to developmental patterning, we have had a long standing interest in understanding how graded BMP signaling establishes cell fates along the dorsal-ventral (DV) axis of Drosophila embryos and how wing vein form along borders between discrete domains in the wing imaginal disc. Our studies on BMPs in DV patterning began with our cloning of the short gastrulation ( sog) gene [1], which encodes a secreted BMP antagonist homologous to vertebrate Chordin [2].  In collaboration with David Kimelman’s group (Univ. Washington), we showed that sog activity was conserved in Xenopus embryos where, like chordin, it could induce a secondary neural axis. Our studies also revealed that Sog acts to prevent BMP signaling from spreading ventrally into the Drosophila neuroectoderm via an autoactivation loop, a process that is also conserved in vertebrate embryos [3], thus helping to reverse the prior paradigm for neural induction as a positive-acting process as originally formulated by Hans Spemann in the early 20th century. We also have studied formation of the Sog protein gradient in the dorsal epidermal region of the embryo which we visualized [4] and analyzed quantitatively in collaboration with Arthur Lander’s group (UCI) [5]. This BMP activity gradient arises as a consequence of an adjacent source/sink configuration wherein ventrally produced Sog diffuses dorsally, where it is degraded by the metalloprotease Tolloid and removed by endocytosis [4]. Most recently, in collaboration with Shannon Fisher’s group (U Penn) and Giorgos Pyrowolakis (Univ. Freiburg) we have been examining the role that BMPs play in patterning the dorsal and lateral regions of the CNS in both flies and zebrafish. In flies, BMPs function as they do during neural induction by repressing expression of the neural identity genes msh and ind (whichdefine dorsal and lateral CNS cell fates, respectively). A twist , however, is that BMPs act in a dose-dependent fashion to repress ind more efficiently than msh, thereby providing a patterning cue [6]. These experiments should shed light on the evolutionary origins of the condensed CNS, which we believe arose once in the ancestors of bilateria [7, 8].
In collaboration with David Kimelman’s group (Univ. Washington), we showed that sog activity was conserved in Xenopus embryos where, like chordin, it could induce a secondary neural axis. Our studies also revealed that Sog acts to prevent BMP signaling from spreading ventrally into the Drosophila neuroectoderm via an autoactivation loop, a process that is also conserved in vertebrate embryos [3], thus helping to reverse the prior paradigm for neural induction as a positive-acting process as originally formulated by Hans Spemann in the early 20th century. We also have studied formation of the Sog protein gradient in the dorsal epidermal region of the embryo which we visualized [4] and analyzed quantitatively in collaboration with Arthur Lander’s group (UCI) [5]. This BMP activity gradient arises as a consequence of an adjacent source/sink configuration wherein ventrally produced Sog diffuses dorsally, where it is degraded by the metalloprotease Tolloid and removed by endocytosis [4]. Most recently, in collaboration with Shannon Fisher’s group (U Penn) and Giorgos Pyrowolakis (Univ. Freiburg) we have been examining the role that BMPs play in patterning the dorsal and lateral regions of the CNS in both flies and zebrafish. In flies, BMPs function as they do during neural induction by repressing expression of the neural identity genes msh and ind (whichdefine dorsal and lateral CNS cell fates, respectively). A twist , however, is that BMPs act in a dose-dependent fashion to repress ind more efficiently than msh, thereby providing a patterning cue [6]. These experiments should shed light on the evolutionary origins of the condensed CNS, which we believe arose once in the ancestors of bilateria [7, 8].

Our studies on wing vein patterning are focused on how primary positional information provided by BMP and Hedgehog morphogen gradients is refined into sharp borders abutting broad gene expression domains to define the positions along which wing vein primordia will develop [9, 10]. Wing veins are rigid hollow fluid conducting channels that function as structural supports for flight and as conduits bringing nutrients to sensory organs in the wing. Although formation of each vein is initiated separately, these linear structures are all induced by the same general mechanism (termed “for-export-only signaling”) in which cells in one domain produce a diffusible factor to which they are unable to respond [10]. The vein inducing signal diffuses into the adjacent cell territory where cells nearest the source of the signal can respond by activating expression of vein specific identity genes such as knirps (for the L2 vein) [11, 12] or abrupt (for the L5 vein) [13]. Most recently we have become interested in the evolution of the gene regulatory networks underlying wing development in dipteral species with different vein patterns than in D. melanogaster.
Analyzing Mechanisms of Human Disease in Flies
With regard to using Drosophila as tool to study human disease [14], we have used this system to identify the Rho-GEF pebble as a candidate gene involved in Angelman syndrome [15], to examine an interaction between g-secretase and peroxiredoxins that may be relevant to Alzheimer’s disease [16], to identify a cooperative interaction between the cell adhesion molecule DSCAM and a collagen subunit (COL6A2) that when over-expressed leads to congenital heart defects in flies and mice and may contribute to heart defects in Down syndrome patients (in collaboration with Geoff Rosenfeld, UCSD, Medicine) [17], and, in collaboration with Michael Karin (UCSD, Pharmacology) to examine mechanisms of metabolic homeostasis relevant to aging [18]. Our current focus is on identifying genetic pathways that are disrupted by pathogenic bacterial and viral factors. For example, in collaboration with Michael Karin and Victor Nizet (UCSD, Pediatrics) we have examined the cell biological effects of anthrax toxins [19-21]. These studies identified the small GTPase Rab11 and its exocyst binding partner Sec15 as targets of the barrier disruptive anthrax toxins Edema Factor (EF) and Lethal Factor (LF) [19].  EF functions by down-regulating expression and activity of Rab11 on late recycling endosomal membranes, and thereby indirectly leads to down-regulation of Sec15, which tethers the endosome to docking sites in the plasma membrane. LF does not affect Rab11, but does inhibit Sec15 expression, and functions cooperatively with EF to disrupt exocyst mediated trafficking and endothelial barrier integrity. We are also conducting a similar analysis on the effect of cholera toxin (Ctx, an ADP ribosylase that activates Gsa proteins thereby constitutively activating endogenous adenylate cyclase to produce elevated levels of cAMP) in intestinal epithelial cells [22]. These studies have revealed that in addition to its classic well characterized role in inducing transcellular Cl- ion secretion via the CFTR channel, Ctx also weakens cell junctions, which may facilitate the accompanying efflux of Na+ counter ions and water via the paracellular route, thus resulting in the profuse salty diarrhea characteristic of cholera. Importantly, the junction disrupting effects of both EF and Ctx can be reversed by over-expressing Rab11, suggesting a new avenue for therapeutic intervention in these diseases. In collaboration with David Broide (UCSD, Medicine) we have also begun to examine whether endocytic recycling plays a role maintaining barrier integrity in airway epithelial cells, and whether defects in this trafficking contribute to asthma pathogenesis. Insights derived from these synergistic studies will have relevance to a broad range of diseases that increase epithelial or endothelial permeability and may lead to therapeutic strategies that selectively counteract such effects by restoring endocytic trafficking and improving barrier function.
EF functions by down-regulating expression and activity of Rab11 on late recycling endosomal membranes, and thereby indirectly leads to down-regulation of Sec15, which tethers the endosome to docking sites in the plasma membrane. LF does not affect Rab11, but does inhibit Sec15 expression, and functions cooperatively with EF to disrupt exocyst mediated trafficking and endothelial barrier integrity. We are also conducting a similar analysis on the effect of cholera toxin (Ctx, an ADP ribosylase that activates Gsa proteins thereby constitutively activating endogenous adenylate cyclase to produce elevated levels of cAMP) in intestinal epithelial cells [22]. These studies have revealed that in addition to its classic well characterized role in inducing transcellular Cl- ion secretion via the CFTR channel, Ctx also weakens cell junctions, which may facilitate the accompanying efflux of Na+ counter ions and water via the paracellular route, thus resulting in the profuse salty diarrhea characteristic of cholera. Importantly, the junction disrupting effects of both EF and Ctx can be reversed by over-expressing Rab11, suggesting a new avenue for therapeutic intervention in these diseases. In collaboration with David Broide (UCSD, Medicine) we have also begun to examine whether endocytic recycling plays a role maintaining barrier integrity in airway epithelial cells, and whether defects in this trafficking contribute to asthma pathogenesis. Insights derived from these synergistic studies will have relevance to a broad range of diseases that increase epithelial or endothelial permeability and may lead to therapeutic strategies that selectively counteract such effects by restoring endocytic trafficking and improving barrier function.
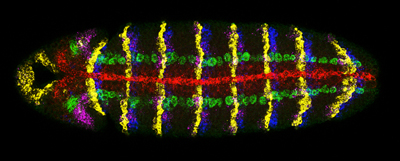
Quantitative Imaging and Genome Analysis
In collaboration with the McGinnis group (UCSD, CDB) and Terry Hwa's lab (UCSD, Physics) we have also developed new methods and quantitative tools to detect multiple gene expression patterns in single cells using multiplex in situ hybridization [23] as well as computational techniques to quantitatively measure gene expression levels [24]. We have applied these to generate testable models of pattern formation in both the developing embryo and wing. For example, most recently, we have proposed that differences in gene length alone can create a transient nested pattern of gene expression to help “seed” morphogen-dependent patterning across a developing field of cells [24]. We have also developed bioinformatic tools for mining and comparing genome data [25, 26].
Select Publications
- Francois, V., M. Solloway, J.W. O'Neill, J. Emery, and E. Bier. Dorsal-ventral patterning of the Drosophila embryo depends on a putative negative growth factor encoded by the short gastrulation gene . 1994. Genes Dev 8: 2602-16.
- Francois, V. and E. Bier. Xenopus chordin and Drosophila short gastrulation genes encode homologous proteins functioning in dorsal-ventral axis formation . 1995. Cell 80: 19-20.
- Biehs, B., V. Francois, and E. Bier. The Drosophila short gastrulation gene prevents Dpp from autoactivating and suppressing neurogenesis in the neuroectoderm . 1996. Genes Dev 10: 2922-34.
- Srinivasan, S., K.E. Rashka, and E. Bier. Creation of a Sog morphogen gradient in the Drosophila embryo . 2 002. Dev Cell 2: 91-101.
- Mizutani, C.M., Q. Nie, F.Y. Wan, Y.T. Zhang, P. Vilmos, R. Sousa-Neves, E. Bier, J.L. Marsh, and A.D. Lander. Formation of the BMP activity gradient in the Drosophila embryo . 2005. Dev Cell 8: 915-24.
- Mizutani, C.M., N. Meyer, H. Roelink, and E. Bier. Threshold-Dependent BMP-Mediated Repression: A Model for a Conserved Mechanism That Patterns the Neuroectoderm . 2006. PLoS Biology 4: e313.
- Mizutani, C.M. and E. Bier. EvoD/Vo: the origins of BMP signalling in the neuroectoderm . 2008. Nat Rev Genet.
- Bier, E. Evolution of development: diversified dorsoventral patterning . 2011. Curr Biol 21: R591-4.
- Biehs, B., M.A. Sturtevant, and E. Bier. Boundaries in the Drosophila wing imaginal disc organize vein-specific genetic programs . 1998. Development 125: 4245-57.
- Bier, E. Drawing lines in the Drosophila wing: initiation of wing vein development . 2000. Curr Opin Genet Dev 10: 393-8.
- Lunde, K., B. Biehs, U. Nauber, and E. Bier. The knirps and knirps-related genes organize development of the second wing vein in Drosophila . 1998. Development 125: 4145-54.
- Lunde, K., J.L. Trimble, A. Guichard, K.A. Guss, U. Nauber, and E. Bier. Activation of the knirps locus links patterning to morphogenesis of the second wing vein in Drosophila . 2003. Development 130: 235-48.
- Cook, O., B. Biehs, and E. Bier. brinker and optomotor-blind act coordinately to initiate development of the L5 wing vein primordium in Drosophila . 2004. Development 131: 2113-24.
- Bier, E. Drosophila, the golden bug, emerges as a tool for human genetics . 2005. Nat Rev Genet 6: 9-23.
- Reiter, L.T., T.N. Seagroves, M. Bowers, and E. Bier. Expression of the Rho-GEF Pbl/ECT2 is regulated by the UBE3A E3 ubiquitin ligase . 2006. Hum Mol Genet 15: 2825-35.
- Wangler, M.F., L.T. Reiter, G. Zimm, J. Trimble-Morgan, J. Wu, and E. Bier. Antioxidant proteins TSA and PAG interact synergistically with Presenilin to modulate Notch signaling in Drosophila . 2011. Protein Cell 2: 554-63.
- Grossman, T.R., A. Gamliel, R.J. Wessells, O. Taghli-Lamallem., K. Jepsen, K. Ocorr, J.R. Korenberg, K.L. Peterson, M.G. Rosenfeld, R. Bodmer, and E. Bier. Over-expression of DSCAM and COL6A2 cooperatively generates congenital heart defects . 2011. PLoS Genet 7: e1002344.
- Lee, J.H., A.V. Budanov, E.J. Park, R. Birse, T.E. Kim, G.A. Perkins, K. Ocorr, M.H. Ellisman, R. Bodmer, E. Bier, and M. Karin. Sestrin as a feedback inhibitor of TOR that prevents age-related pathologies . 2010. Science 327: 1223-8.
- Guichard, A., S.M. McGillivray, B. Cruz-Moreno, N.M. van Sorge, V. Nizet, and E. Bier. Anthrax toxins cooperatively inhibit endocytic recycling by the Rab11/Sec15 exocyst . 2010. Nature 467: 854-8.
- Guichard, A., V. Nizet, and E. Bier. New insights into the biological effects of anthrax toxins: linking cellular to organismal responses . 2012. Microbes Infect 14: 97-118.
- Guichard, A., J.M. Park, B. Cruz-Moreno, M. Karin, and E. Bier. Anthrax lethal factor and edema factor act on conserved targets in Drosophila . 2006. Proc Natl Acad Sci U S A 103: 3244-9.
- Guichard, A., B. Cruz-Moreno, B. Aguilar, J. Kuang, A.A. Kurkciyan, Z. Wang, S. Hang, G.P. Pineton de Chambrun, D.F. McCole, P. Watnick, N.M. van Sorge, V. Nizet, and E. Bier. (2013). Cholera toxin disrupts barrier function by inhibiting exocyst-mediated trafficking of host proteins to intestinal cell junctions. Cell Host & Microbe 14, 294-305.
- Kosman, D., C.M. Mizutani, D. Lemons, W.G. Cox, W. McGinnis, and E. Bier. Multiplex detection of RNA expression in Drosophila embryos . 2004. Science 305: 846.
- McHale, P., C.M. Mizutani, D. Kosman, D.L. MacKay, M. Belu, A. Hermann, W. McGinnis, E. Bier, and T. Hwa. Gene length may contribute to graded transcriptional responses in the Drosophila embryo . 2011. Dev Biol 360: 230-40.
- Reiter, L.T., L. Potocki, S. Chien, M. Gribskov, and E. Bier. A systematic analysis of human disease-associated gene sequences in Drosophila melanogaster . 2001. Genome Res 11: 1114
- Do, L.H., F.F. Esteves, H.J. Karten, and E. Bier. Booly: a new data integration platform . 2010. BMC Bioinformatics 11: 513.
Biography
Ethan Bier received his Ph.D. from Harvard University. He has received the Sloan Foundation Award, the McKnight Award for Neuroscience, the Basil O'Connor Award, and the American Cancer Society Junior Faculty Award.
