Li-Fan Lu
Research
During the last decade a unique mechanism of negative regulation of immune responses and inflammation by a dedicated population of so-called regulatory T (Treg) cells has become the focus of intensive investigation. Expression of the transcription factor Foxp3, a hallmark of Treg cells, is necessary for the establishment and maintenance of a distinct transcriptional program that determines metabolic, signaling, and effector features (e.g., suppressive function), distinguishing these cells from other T cell lineages. The long-term goals of our research are to identify the molecular determinants of differentiation and function of Treg cells and to ultimately gain insight into the complex biology of these cells, including the maintenance of immunological tolerance to "self" and the regulation of immune responses to pathogens, commensals, and tumors.
1. Identification of miRNAs crucial for mediating Treg cell biology in healthy and diseased mice
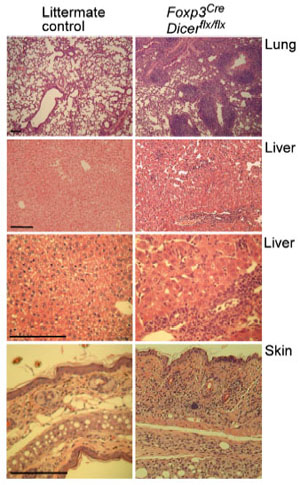
Several thousands genes are differentially expressed in Foxp3+ Treg cells, including a subset of non-coding small regulatory microRNA (miRNA), some of which are directly regulated by Foxp3 suggesting a role for miRNA-mediated regulation of gene expression in Treg cell differentiation, maintenance or function. Catastrophic consequences and the development of complex disease phenotypes in mice harboring Treg cells devoid of miRNA suggested miRNAs control multiple facets of Treg cell biology (Fig.1). Furthermore, our subsequent studies in miR-155 and miR-146a have demonstrated that individual miRNA could affect distinct aspect of Treg cell biology. By employing genetic, biochemical, immunological approaches and whole animal experimentation, we aim to identify miRNA expression profiles in Treg cells isolated from different tissues in unmanipulated mice as well as in mice suffering different pathologies. Moreover, we are interested in examining how miRNAs themselves are regulated in response to different environmental cues. Ultimately, we hope to elucidate how such regulation of miRNA expression impacts their respective targets and how the dynamic miRNA-dependent regulation enforces Treg cell functionality in diverse disease settings.
2. Elucidating molecular and cellular mechanisms by which Treg cells control type I inflammation
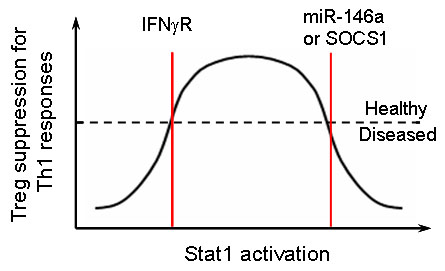
Recent studies revealed that the expression of Th lineage-specific transcription factors in Treg cells is crucial for Treg cell-mediated control of corresponding immune responses in particular tissue and inflammatory settings. Our analysis of the role of miR-146a and SOCS1 in Treg cell-dependent Th1 regulation have further extended this concept and demonstrated that both lack of Stat1 and unrestrained Stat1 activation in Treg cells would result in a similar breakdown of immunologic tolerance (Fig.2). These results suggest that Treg cell-mediated suppression of a given Th-specific immunopathology is depending upon an optimal range of activation in corresponding Th-lineage specific transcription factors. One fundamental question that remained is how Treg cells control Th1 responses. At the molecular level, we are interested in identifying important effector molecules and signalling pathways underlying Treg cell-mediated Th1 regulation. At the cellular level, we hope to understand the complex interplay between Treg cells, effector T cells and other immune cells in the surrounding environment specifically in a type I inflammatory setting. Moreover, IFNγ-mediated Th1 responses and Treg cells have been associated with inflammation in many clinical settings such as infection (e.g., T. gondii), autoimmune diseases (e.g., multiple sclerosis and rheumatoid arthritis), tumors, and organ transplantation. Through our study, we hope to establish the foundation for the future development of means to manipulating Treg and T effector cell function for better treatment in human patients.
Select Publications
- Dong, J.*, Warner, L.M.*, Lin, L.L., Chen, M.C., O'Connell, R.M. and Lu, L.F. (*these authors contributed equally) 2021. MiR-155 promotes Treg cell development by safeguarding medullary thymic epithelial cell maturation. J Exp Med. 218(2): e20192423.
- Wu, C.J., Cho, S., Huang, H.Y., Lu, C.H., Russ, J., Cruz, L.O., da Cunha, F.F., Chen, M.C., Lin, L.L., Warner, L.M., Liao, H.K., Utzschneider, D.T., Quon, S., Berner, J., Camara, N.O.S., Zehn, D., Izpisua Belmonte, J.C., Chen, L.C., Huang, S.F., Kuo, M.L. and Lu, L.F. 2019. MiR-23~27~24-mediated control of humoral immunity reveals a TOX-driven regulatory circuit in follicular helper T cell differentiation. Sci Adv. 5(12): eaaw1715. PMCID: PMC6905868
- Dong, J., Tai, J.W. and Lu, L.F. 2019. MicroRNA-microbiota Interaction in Gut Homeostasis and Colorectal Cancer. Trends Cancer 5(11): 666-669.
- Cho, S.*, Lee, H.M.*, Yu, I.S., Choi, Y.S., Huang, H.Y., Hashemifar, S.S., Lin, L.L., Chen, M.C., Afanasiev, N.D., Khan, A.A., Lin, S.W., Rudensky, A.Y., Crotty, S. and Lu, L.F. (*these authors contributed equally) 2018. Differential cell-intrinsic regulations of germinal center B and T cells by miR-146a and miR-146b. Nat Commun. 9(1):2757. PMCID: PMC6048122
- Becher, B., Waisman, A. and Lu, L.F. 2018. Conditional Gene-Targeting in Mice: Problems and Solutions. Immunity 48(5):835-836.
- Cruz, L.O., Hashemifar, S.S., Wu, C.J., Cho, S., Nguyen, D.T., Lin, L.L., Khan, A.A. and Lu, L.F. 2017. Excessive expression of miR-27 impairs regulatory T cell-mediated immunological tolerance. J. Clin. Invest. 127(2):530-42. PMCID: PMC5272185
- Cho, S.*, Wu, C.J.*, Yasuda, T.*, Cruz, L.O., Khan, A.A., Lin, L.L., Nguyen, D.T., Miller, M., Lee, H.M., Kuo, M.L., Broide, D.H., Rajewsky, K., Rudensky, A.Y. and Lu, L.F. (*these authors contributed equally) 2016. MiR-23~27~24 clusters control effector T cell differentiation and function. J. Exp. Med. 213(2):235-49. PMCID: PMC4749926
- Lu, L.F.*, Gasteiger, G.*, Yu, I.S., Chaudhry, A., Hsin, J.P., Lu, Y., Bos, P.D., Lin, L.L., Zawislak. C.L., Cho, S., Sun, J.C., Leslie, C.S., Lin, S.W. and Rudensky, A.Y. (*these authors contributed equally) 2015. A single miRNA-mRNA interaction affects the immune response in a context- and cell-type-specific manner. Immunity 43(1):52-64. PMCID: PMC4529747
- Lee, H.M., Fleige, A., Forman, R., Cho, S., Khan, A.A., Lin, L.L., Nguyen, D.T., O’Hara-Hall, A., Yin, Z., Hunter, C.A., Muller, W. and Lu, L.F. 2015. IFNγ signaling endows DCs with the capacity to control type I inflammation during parasitic infection through promoting T-bet+ regulatory T cells. PLOS Pathog. 11(2):e1004635. PMCID: PMC4450074
- Josefowicz, S.Z.*, Lu, L.F.* and Rudensky, A.Y. (*these authors contributed equally) 2012. Regulatory T cells: mechanisms of differentiation and function. Annu. Rev. Immunol. 30:531-64
- Lu, L.F., Boldin, M.P., Chaudhry, A., Lin, L.L., Taganov, K.D., Hanada, T., Yoshimura, A., Baltimore, D. and Rudensky, A.Y. 2010. Function of miR-146a in controlling Treg cell-mediated regulation of Th1 responses. Cell 142(6): 914-29. PMCID: PMC3049116
- Lu, L.F. and Rudensky A. 2009. Molecular orchestration of differentiation and function of regulatory T cells. Genes Dev. 23(11): 1270-1282. PMCID: PMC2763538
- Lu, L.F.*, Thai, T.H.*, Calado, D.P.*, Chaudhry, A., Kubo, M., Tanaka, K., Loeb, G.B., Lee, H., Yoshimura, A., Rajewsy, K. and Rudensky, A.Y. (*these authors contributed equally) 2009. Foxp3-dependent microRNA155 confers competitive fitness to regulatory T cells through targeting SOCS1 protein. Immunity 30(1):80-91. PMCID:PMC2654249
- Liston, A.*, Lu, L.F.*, Carroll D.O., Tarakhovsky, A. and Rudensky, A.Y. (*these authors contributed equally) 2008. Dicer-dependent microRNA pathway safeguards regulatory T cell function. J. Exp. Med. 205(9):1993-2004. PMCID: PMC2526195
- Lu, L.F., Lind, E.F., Gondek, D.C., Bennett, K.A., Gleeson, M.W., Pino-Lagos, K., Scott, Z.A., Coyle, A.J., Reed, J.L., Van Snick, J, Strom, T.B., Zheng, X.X. and Noelle R.J. 2006. Mast cells as essential intermediaries in regulatory T cell tolerance. Nature 442 (7106): 997-1002.
Biography
Li-Fan Lu received his Ph.D. in Immunology from Dartmouth College. He did his postdoctoral training first at the University of Washington and then at the Memorial Sloan-Kettering Cancer Center where he was the Leukemia and Lymphoma Society Fellow. In 2011, he received the K99/R00 Career Development Award through NIAID.
